-
 The Therapeutic Mechanisms of Mesenchymal Stem Cells in MS—A Review Focusing on Neuroprotective Properties
The Therapeutic Mechanisms of Mesenchymal Stem Cells in MS—A Review Focusing on Neuroprotective Properties -
 The Bee Gut Microbiota: Bridging Infective Agents Potential in the One Health Context
The Bee Gut Microbiota: Bridging Infective Agents Potential in the One Health Context -
 The Role of Extracellular Vesicles in Atopic Dermatitis
The Role of Extracellular Vesicles in Atopic Dermatitis -
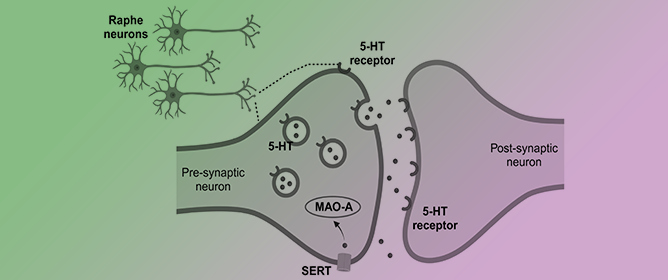 Revisiting the Role of Serotonin in Sleep-Disordered Breathing
Revisiting the Role of Serotonin in Sleep-Disordered Breathing -
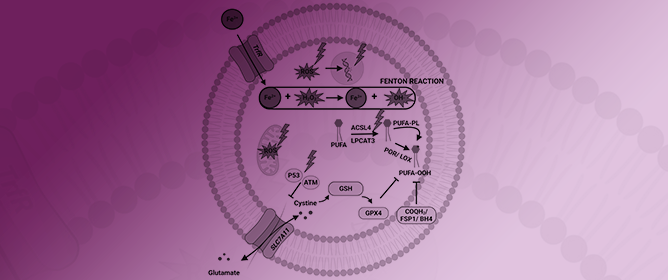 Ferroptosis: Frenemy of Radiotherapy
Ferroptosis: Frenemy of Radiotherapy
Journal Description
International Journal of Molecular Sciences
International Journal of Molecular Sciences
is an international, peer-reviewed, open access journal providing an advanced forum for biochemistry, molecular and cell biology, molecular biophysics, molecular medicine, and all aspects of molecular research in chemistry, and is published semimonthly online by MDPI. The Australian Society of Plant Scientists (ASPS), Epigenetics Society, European Calcium Society (ECS), European Chitin Society (EUCHIS), Spanish Society for Cell Biology (SEBC) and others are affiliated with IJMS and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, MEDLINE, Embase, CAPlus / SciFinder, and other databases.
- Journal Rank: JCR - Q1 (Biochemistry & Molecular Biology) / CiteScore - Q1 (Inorganic Chemistry)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.3 days after submission; acceptance to publication is undertaken in 2.6 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about the IJMS.
- Companion journals for IJMS include: Biophysica, Obesities, Stresses and Lymphatics.
Impact Factor:
5.6 (2022);
5-Year Impact Factor:
6.2 (2022)
Latest Articles
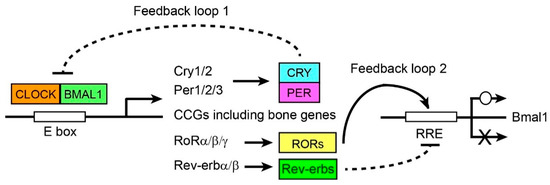
Circadian Regulation of Bone Remodeling
Int. J. Mol. Sci. 2024, 25(9), 4717; https://doi.org/10.3390/ijms25094717 (registering DOI) - 26 Apr 2024
Abstract
Adult bones are continuously remodeled by the balance between bone resorption by osteoclasts and subsequent bone formation by osteoblasts. Many studies have provided molecular evidence that bone remodeling is under the control of circadian rhythms. Circadian fluctuations have been reported in the serum
[...] Read more.
Adult bones are continuously remodeled by the balance between bone resorption by osteoclasts and subsequent bone formation by osteoblasts. Many studies have provided molecular evidence that bone remodeling is under the control of circadian rhythms. Circadian fluctuations have been reported in the serum and urine levels of bone turnover markers, such as digested collagen fragments and bone alkaline phosphatase. Additionally, the expressions of over a quarter of all transcripts in bones show circadian rhythmicity, including the genes encoding master transcription factors for osteoblastogenesis and osteoclastogenesis, osteogenic cytokines, and signaling pathway proteins. Serum levels of calcium, phosphate, parathyroid hormone, and calcitonin also display circadian rhythmicity. Finally, osteoblast- and osteoclast-specific knockout mice targeting the core circadian regulator gene Bmal1 show disrupted bone remodeling, although the results have not always been consistent. Despite these studies, however, establishing a direct link between circadian rhythms and bone remodeling in vivo remains a major challenge. It is nearly impossible to repeatedly collect bone materials from human subjects while following circadian changes. In addition, the differences in circadian gene regulation between diurnal humans and nocturnal mice, the main model organism, remain unclear. Filling the knowledge gap in the circadian regulation of bone remodeling could reveal novel regulatory mechanisms underlying many bone disorders including osteoporosis, genetic diseases, and fracture healing. This is also an important question for the basic understanding of how cell differentiation progresses under the influence of cyclically fluctuating environments.
Full article
(This article belongs to the Special Issue Advances in Bone Growth, Development and Metabolism)
►
Show Figures
Open AccessArticle
An Explainable Deep Learning Classifier of Bovine Mastitis Based on Whole-Genome Sequence Data—Circumventing the p >> n Problem
by
Krzysztof Kotlarz, Magda Mielczarek, Przemysław Biecek, Katarzyna Wojdak-Maksymiec, Tomasz Suchocki, Piotr Topolski, Wojciech Jagusiak and Joanna Szyda
Int. J. Mol. Sci. 2024, 25(9), 4715; https://doi.org/10.3390/ijms25094715 (registering DOI) - 26 Apr 2024
Abstract
The serious drawback underlying the biological annotation of whole-genome sequence data is the p >> n problem, which means that the number of polymorphic variants (p) is much larger than the number of available phenotypic records (n). We propose a way to circumvent
[...] Read more.
The serious drawback underlying the biological annotation of whole-genome sequence data is the p >> n problem, which means that the number of polymorphic variants (p) is much larger than the number of available phenotypic records (n). We propose a way to circumvent the problem by combining a LASSO logistic regression with deep learning to classify cows as susceptible or resistant to mastitis, based on single nucleotide polymorphism (SNP) genotypes. Among several architectures, the one with 204,642 SNPs was selected as the best. This architecture was composed of two layers with, respectively, 7 and 46 units per layer implementing respective drop-out rates of 0.210 and 0.358. The classification of the test data resulted in AUC = 0.750, accuracy = 0.650, sensitivity = 0.600, and specificity = 0.700. Significant SNPs were selected based on the SHapley Additive exPlanation (SHAP). As a final result, one GO term related to the biological process and thirteen GO terms related to molecular function were significantly enriched in the gene set that corresponded to the significant SNPs. Our findings revealed that the optimal approach can correctly predict susceptibility or resistance status for approximately 65% of cows. Genes marked by the most significant SNPs are related to the immune response and protein synthesis.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
►▼
Show Figures
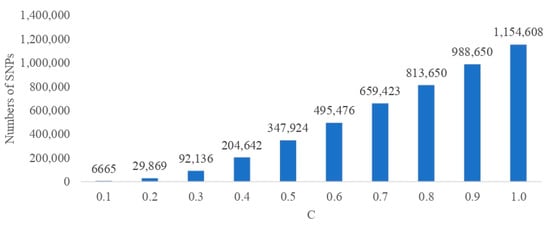
Figure 1
Open AccessArticle
Cytotoxicity of Quantum Dots in Receptor-Mediated Endocytic and Pinocytic Pathways in Yeast
by
Onyinye Okafor and Kyoungtae Kim
Int. J. Mol. Sci. 2024, 25(9), 4714; https://doi.org/10.3390/ijms25094714 (registering DOI) - 26 Apr 2024
Abstract
Despite the promising applications of the use of quantum dots (QDs) in the biomedical field, the long-lasting effects of QDs on the cell remain poorly understood. To comprehend the mechanisms underlying the toxic effects of QDs in yeast, we characterized defects associated with
[...] Read more.
Despite the promising applications of the use of quantum dots (QDs) in the biomedical field, the long-lasting effects of QDs on the cell remain poorly understood. To comprehend the mechanisms underlying the toxic effects of QDs in yeast, we characterized defects associated with receptor-mediated endocytosis (RME) as well as pinocytosis using Saccharomyces cerevisiae as a model in the presence of cadmium selenide/zinc sulfide (CdSe/ZnS) QDs. Our findings revealed that QDs led to an inefficient RME at the early, intermediate, and late stages of endocytic patch maturation at the endocytic site, with the prolonged lifespan of GFP fused yeast fimbrin (Sac6-GFP), a late marker of endocytosis. The transit of FM1-43, a lipophilic dye from the plasma membrane to the vacuole, was severely retarded in the presence of QDs. Finally, QDs caused an accumulation of monomeric red fluorescent protein fused carbamoyl phosphate synthetase 1 (mRFP-Cps1), a vacuolar lumen marker in the vacuole. In summary, the present study provides novel insights into the possible impact of CdSe/ZnS QDs on the endocytic machinery, enabling a deeper comprehension of QD toxicity.
Full article
(This article belongs to the Section Molecular Nanoscience)
►▼
Show Figures
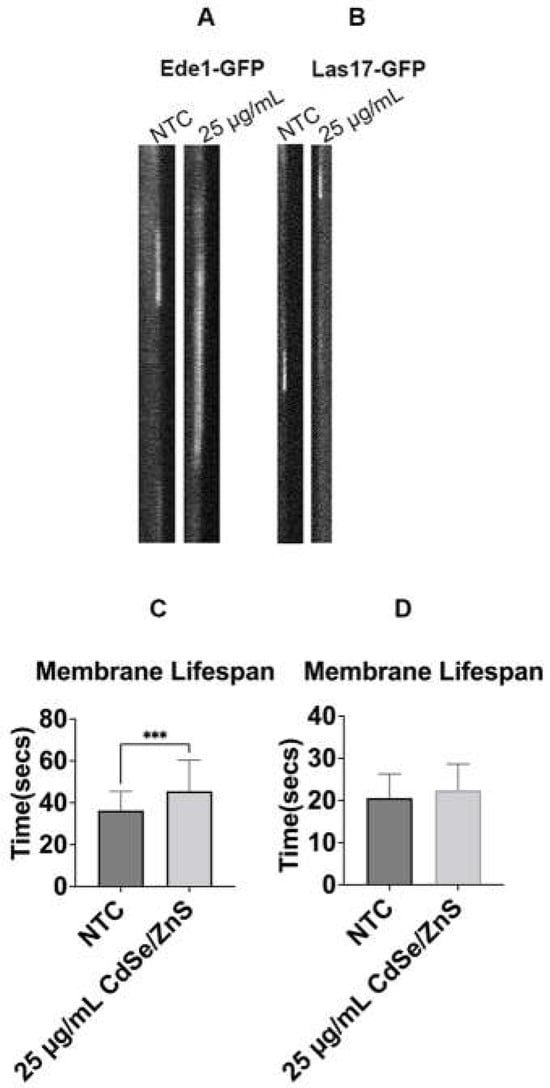
Figure 1
Open AccessArticle
(2,6-Dimethylphenyl)arsonic Acid Induces Apoptosis through the Mitochondrial Pathway, Downregulates XIAP, and Overcomes Multidrug Resistance to Cytostatic Drugs in Leukemia and Lymphoma Cells In Vitro
by
Nathalie Wilke, Corazon Frias, Albrecht Berkessel and Aram Prokop
Int. J. Mol. Sci. 2024, 25(9), 4713; https://doi.org/10.3390/ijms25094713 (registering DOI) - 26 Apr 2024
Abstract
Cancer treatment is greatly challenged by drug resistance, highlighting the need for novel drug discoveries. Here, we investigated novel organoarsenic compounds regarding their resistance-breaking and apoptosis-inducing properties in leukemia and lymphoma. Notably, the compound (2,6-dimethylphenyl)arsonic acid (As2) demonstrated significant inhibition of cell proliferation
[...] Read more.
Cancer treatment is greatly challenged by drug resistance, highlighting the need for novel drug discoveries. Here, we investigated novel organoarsenic compounds regarding their resistance-breaking and apoptosis-inducing properties in leukemia and lymphoma. Notably, the compound (2,6-dimethylphenyl)arsonic acid (As2) demonstrated significant inhibition of cell proliferation and induction of apoptosis in leukemia and lymphoma cells while sparing healthy leukocytes. As2 reached half of its maximum activity (AC50) against leukemia cells at around 6.3 µM. Further experiments showed that As2 overcomes multidrug resistance and sensitizes drug-resistant leukemia and lymphoma cell lines to treatments with the common cytostatic drugs vincristine, daunorubicin, and cytarabine at low micromolar concentrations. Mechanistic investigations of As2-mediated apoptosis involving FADD (FAS-associated death domain)-deficient or Smac (second mitochondria-derived activator of caspases)/DIABLO (direct IAP binding protein with low pI)-overexpressing cell lines, western blot analysis of caspase-9 cleavage, and measurements of mitochondrial membrane integrity identified the mitochondrial apoptosis pathway as the main mode of action. Downregulation of XIAP (x-linked inhibitor of apoptosis protein) and apoptosis induction independent of Bcl-2 (B-cell lymphoma 2) and caspase-3 expression levels suggest the activation of additional apoptosis-promoting mechanisms. Due to the selective apoptosis induction, the synergistic effects with common anti-cancer drugs, and the ability to overcome multidrug resistance in vitro, As2 represents a promising candidate for further preclinical investigations with respect to refractory malignancies.
Full article
(This article belongs to the Special Issue New Agents and Novel Drugs Use for the Oncological Diseases Treatment)
►▼
Show Figures
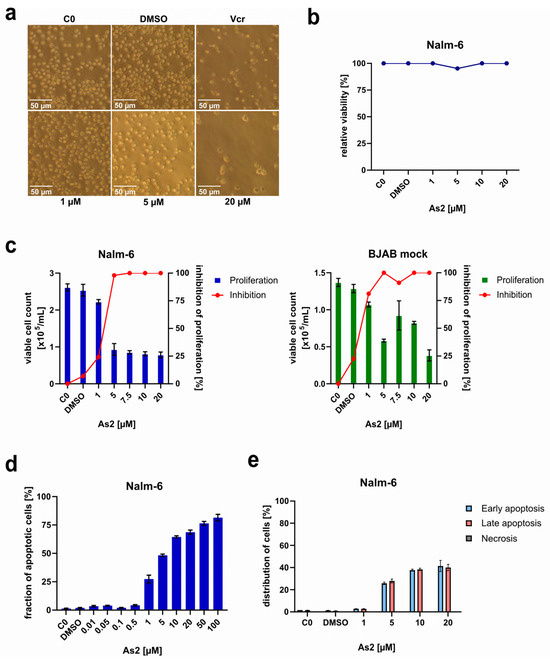
Figure 1
Open AccessArticle
Dual Deletion of Keap1 and Rbpjκ Genes in Liver Leads to Hepatomegaly and Hypercholesterolemia
by
Nobunao Wakabayashi, Yoko Yagishita, Tanvi Joshi and Thomas W. Kensler
Int. J. Mol. Sci. 2024, 25(9), 4712; https://doi.org/10.3390/ijms25094712 (registering DOI) - 26 Apr 2024
Abstract
The hepatic deletion of Rbpjκ (RbpjF/F::AlbCre) in the mouse leads to exhibition of the Alagille syndrome phenotype during early postnatal liver development with hyperlipidemia and cholestasis due to attenuated disruption of NOTCH signaling. Given the roles of NRF2 signaling
[...] Read more.
The hepatic deletion of Rbpjκ (RbpjF/F::AlbCre) in the mouse leads to exhibition of the Alagille syndrome phenotype during early postnatal liver development with hyperlipidemia and cholestasis due to attenuated disruption of NOTCH signaling. Given the roles of NRF2 signaling in the regulation of lipid metabolism and bile ductal formation, it was anticipated that these symptoms could be alleviated by enhancing NRF2 signaling in the RbpjF/F::AlbCre mouse by hepatic deletion of Keap1 in compound Keap1F/F::RbpjF/F::AlbCre mice. Unexpectedly, these mice developed higher hepatic and plasma cholesterol levels with more severe cholestatic liver damage during the pre-weaning period than in the RbpjF/F::AlbCre mice. In addition, hypercholesterolemia and hepatic damage were sustained throughout the growth period unlike in the RbpjF/F::AlbCre mouse. These enhanced abnormalities in lipid metabolism appear to be due to NRF2-dependent changes in gene expression related to cholesterol synthetic and subsequent bile acid production pathways. Notably, the hepatic expression of Cyp1A7 and Abcb11 genes involved in bile acid homeostasis was significantly reduced in Keap1F/F::RbpjF/F::AlbCre compared to RbpjF/F::AlbCre mice. The accumulation of liver cholesterol and the weakened capacity for bile excretion during the 3 pre-weaning weeks in the Keap1F/F::RbpjF/F::AlbCre mice may aggravate hepatocellular damage level caused by both excessive cholesterol and residual bile acid toxicity in hepatocytes. These results indicate that a tuned balance of NOTCH and NRF2 signaling is of biological importance for early liver development after birth.
Full article
(This article belongs to the Special Issue The Role of NRF2 in Health and Disease)
►▼
Show Figures
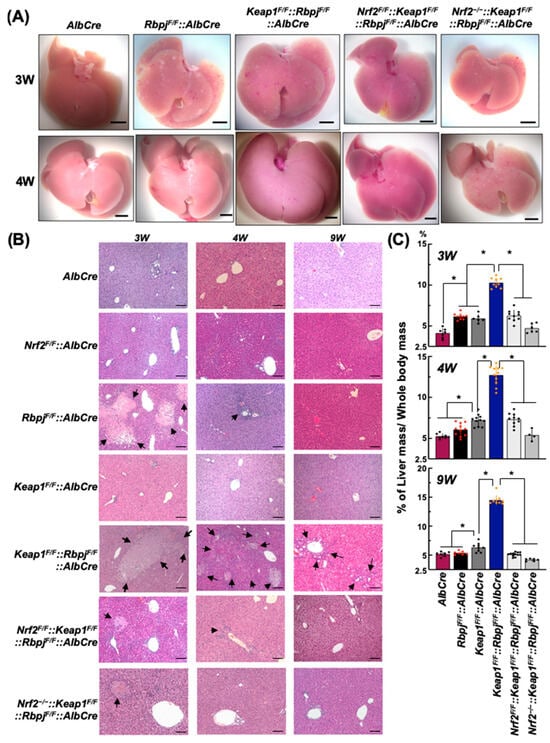
Figure 1
Open AccessArticle
Functional Characterization of the MeSSIII-1 Gene and Its Promoter from Cassava
by
Xiao-Hua Lu, Ya-Jie Wang, Xing-Hou Zhen, Hui Yu, Mu Pan, Dong-Qing Fu, Rui-Mei Li, Jiao Liu, Hai-Yan Luo, Xin-Wen Hu, Yuan Yao and Jian-Chun Guo
Int. J. Mol. Sci. 2024, 25(9), 4711; https://doi.org/10.3390/ijms25094711 (registering DOI) - 26 Apr 2024
Abstract
Soluble starch synthases (SSs) play important roles in the synthesis of cassava starch. However, the expression characteristics of the cassava SSs genes have not been elucidated. In this study, the MeSSIII-1 gene and its promoter, from SC8 cassava cultivars, were respectively isolated by
[...] Read more.
Soluble starch synthases (SSs) play important roles in the synthesis of cassava starch. However, the expression characteristics of the cassava SSs genes have not been elucidated. In this study, the MeSSIII-1 gene and its promoter, from SC8 cassava cultivars, were respectively isolated by PCR amplification. MeSSIII-1 protein was localized to the chloroplasts. qRT-PCR analysis revealed that the MeSSIII-1 gene was expressed in almost all tissues tested, and the expression in mature leaves was 18.9 times more than that in tuber roots. MeSSIII-1 expression was induced by methyljasmonate (MeJA), abscisic acid (ABA), and ethylene (ET) hormones in cassava. MeSSIII-1 expression patterns were further confirmed in proMeSSIII-1 transgenic cassava. The promoter deletion analysis showed that the −264 bp to −1 bp MeSSIII-1 promoter has basal activity. The range from −1228 bp to −987 bp and −488 bp to −264 bp significantly enhance promoter activity. The regions from −987 bp to −747 bp and −747 bp to −488 bp have repressive activity. These findings will provide an important reference for research on the potential function and transcriptional regulation mechanisms of the MeSSIII-1 gene and for further in-depth exploration of the regulatory network of its internal functional elements.
Full article
(This article belongs to the Section Molecular Plant Sciences)
►▼
Show Figures
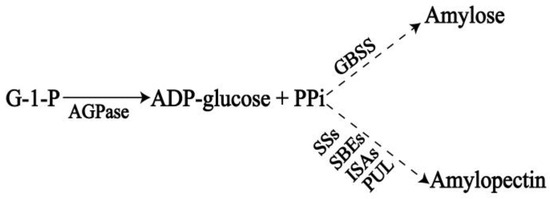
Figure 1
Open AccessReview
Current Perspectives of Mitochondria in Sepsis-Induced Cardiomyopathy
by
Tatsuki Kuroshima, Satoshi Kawaguchi and Motoi Okada
Int. J. Mol. Sci. 2024, 25(9), 4710; https://doi.org/10.3390/ijms25094710 (registering DOI) - 26 Apr 2024
Abstract
Sepsis-induced cardiomyopathy (SICM) is one of the leading indicators for poor prognosis associated with sepsis. Despite its reversibility, prognosis varies widely among patients. Mitochondria play a key role in cellular energy production by generating adenosine triphosphate (ATP), which is vital for myocardial energy
[...] Read more.
Sepsis-induced cardiomyopathy (SICM) is one of the leading indicators for poor prognosis associated with sepsis. Despite its reversibility, prognosis varies widely among patients. Mitochondria play a key role in cellular energy production by generating adenosine triphosphate (ATP), which is vital for myocardial energy metabolism. Over recent years, mounting evidence suggests that severe sepsis not only triggers mitochondrial structural abnormalities such as apoptosis, incomplete autophagy, and mitophagy in cardiomyocytes but also compromises their function, leading to ATP depletion. This metabolic disruption is recognized as a significant contributor to SICM, yet effective treatment options remain elusive. Sepsis cannot be effectively treated with inotropic drugs in failing myocardium due to excessive inflammatory factors that blunt β-adrenergic receptors. This review will share the recent knowledge on myocardial cell death in sepsis and its molecular mechanisms, focusing on the role of mitochondria as an important metabolic regulator of SICM, and discuss the potential for developing therapies for sepsis-induced myocardial injury.
Full article
(This article belongs to the Special Issue Understanding Sepsis: Pathophysiology, Diagnostics and Early Intervention)
►▼
Show Figures
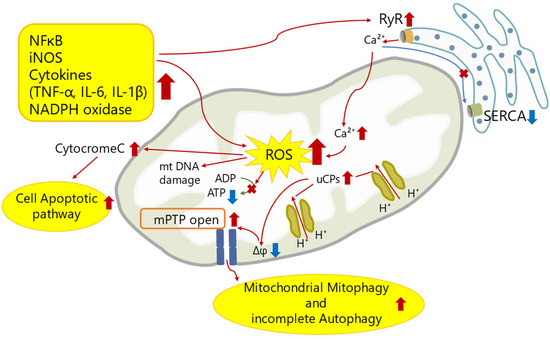
Figure 1
Open AccessEditorial
Special Issue ‘Advances in Neurodegenerative Diseases Research and Therapy 2.0’
by
Sumonto Mitra
Int. J. Mol. Sci. 2024, 25(9), 4709; https://doi.org/10.3390/ijms25094709 (registering DOI) - 26 Apr 2024
Abstract
Neurodegenerative disorders (NDs) and the development of various therapeutic strategies to combat them have received increased attention in recent decades [...]
Full article
(This article belongs to the Special Issue Advances in Neurodegenerative Diseases Research and Therapy 2.0)
Open AccessCase Report
Uncovering the Unseen: Bordetella hinzii Emerges in a Lung Transplant Recipient
by
Damiana-Maria Vulturar, Benoît Pilmis, Claire Rouzaud, Anne Gigandon, Gaëlle Dauriat, Séverine Feuillet-Soummer, Liviu-Stefan Moaca, Elie Fadel, Olaf Mercier, Dominique Fabre, Olivier Lortholary and Jérôme Le Pavec
Int. J. Mol. Sci. 2024, 25(9), 4708; https://doi.org/10.3390/ijms25094708 (registering DOI) - 26 Apr 2024
Abstract
Bordetella hinzii (B. hinzii), a Gram-negative bacillus commonly associated with respiratory infections in animals, has garnered attention for its sporadic cases in humans, particularly in immunocompromised individuals. Despite its opportunistic nature, there remains limited understanding regarding its pathogenicity, diagnostic challenges, and optimal
[...] Read more.
Bordetella hinzii (B. hinzii), a Gram-negative bacillus commonly associated with respiratory infections in animals, has garnered attention for its sporadic cases in humans, particularly in immunocompromised individuals. Despite its opportunistic nature, there remains limited understanding regarding its pathogenicity, diagnostic challenges, and optimal treatment strategies, especially in the context of immunosuppression. Herein, we present the first documented case of acute bronchitis caused by B. hinzii in an immunocompromised patient following double-lung transplantation. The patient, a former smoker with sarcoidosis stage IV, underwent transplant surgery and subsequently developed a febrile episode, leading to the identification of B. hinzii in broncho-alveolar lavage samples. Antimicrobial susceptibility testing revealed resistance to multiple antibiotics, necessitating tailored treatment adjustments. Our case underscores the importance of heightened awareness among clinicians regarding B. hinzii infections and the imperative for further research to elucidate its epidemiology and optimal management strategies, particularly in immunocompromised populations.
Full article
(This article belongs to the Special Issue Immunology of Infectious Disease and Transplantation: A Symbiotic Dance)
►▼
Show Figures
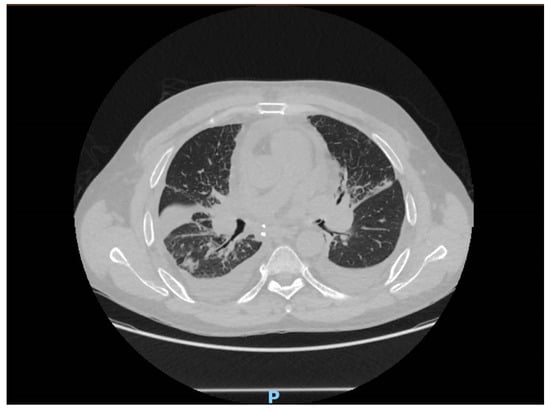
Figure 1
Open AccessArticle
Functional and Genetic Analyses Unveil the Implication of CDC27 in Hemifacial Microsomia
by
Wenjie Song, Xin Xia, Yue Fan, Bo Zhang and Xiaowei Chen
Int. J. Mol. Sci. 2024, 25(9), 4707; https://doi.org/10.3390/ijms25094707 (registering DOI) - 26 Apr 2024
Abstract
Hemifacial microsomia (HFM) is a rare congenital genetic syndrome primarily affecting the first and second pharyngeal arches, leading to defects in the mandible, external ear, and middle ear. The pathogenic genes remain largely unidentified. Whole-exome sequencing (WES) was conducted on 12 HFM probands
[...] Read more.
Hemifacial microsomia (HFM) is a rare congenital genetic syndrome primarily affecting the first and second pharyngeal arches, leading to defects in the mandible, external ear, and middle ear. The pathogenic genes remain largely unidentified. Whole-exome sequencing (WES) was conducted on 12 HFM probands and their unaffected biological parents. Predictive structural analysis of the target gene was conducted using PSIPRED (v3.3) and SWISS-MODEL, while STRING facilitated protein-to-protein interaction predictions. CRISPR/Cas9 was applied for gene knockout in zebrafish. In situ hybridization (ISH) was employed to examine the spatiotemporal expression of the target gene and neural crest cell (NCC) markers. Immunofluorescence with PH3 and TUNEL assays were used to assess cell proliferation and apoptosis. RNA sequencing was performed on mutant and control embryos, with rescue experiments involving target mRNA injections and specific gene knockouts. CDC27 was identified as a novel candidate gene for HFM, with four nonsynonymous de novo variants detected in three unrelated probands. Structural predictions indicated significant alterations in the secondary and tertiary structures of CDC27. cdc27 knockout in zebrafish resulted in craniofacial malformation, spine deformity, and cardiac edema, mirroring typical HFM phenotypes. Abnormalities in somatic cell apoptosis, reduced NCC proliferation in pharyngeal arches, and chondrocyte differentiation issues were observed in cdc27−/− mutants. cdc27 mRNA injections and cdkn1a or tp53 knockout significantly rescued pharyngeal arch cartilage dysplasia, while sox9a mRNA administration partially restored the defective phenotypes. Our findings suggest a functional link between CDC27 and HFM, primarily through the inhibition of CNCC proliferation and disruption of pharyngeal chondrocyte differentiation.
Full article
(This article belongs to the Special Issue Zebrafish: A Powerful Model for Genetics and Genomics 3.0)
►▼
Show Figures
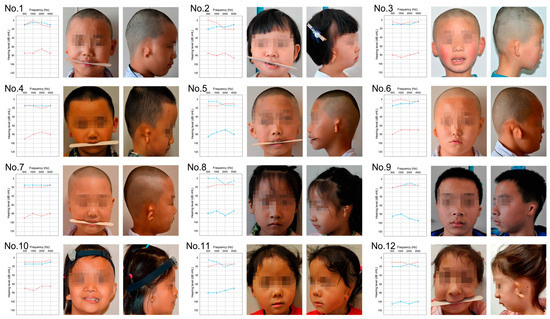
Figure 1
Open AccessArticle
Implementation of FRET Spectrometry Using Temporally Resolved Fluorescence: A Feasibility Study
by
Justin Trujillo, Aliyah S. Khan, Dhruba P. Adhikari, Michael R. Stoneman, Jenu V. Chacko, Kevin W. Eliceiri and Valerica Raicu
Int. J. Mol. Sci. 2024, 25(9), 4706; https://doi.org/10.3390/ijms25094706 (registering DOI) - 26 Apr 2024
Abstract
Förster resonance energy transfer (FRET) spectrometry is a method for determining the quaternary structure of protein oligomers from distributions of FRET efficiencies that are drawn from pixels of fluorescence images of cells expressing the proteins of interest. FRET spectrometry protocols currently rely on
[...] Read more.
Förster resonance energy transfer (FRET) spectrometry is a method for determining the quaternary structure of protein oligomers from distributions of FRET efficiencies that are drawn from pixels of fluorescence images of cells expressing the proteins of interest. FRET spectrometry protocols currently rely on obtaining spectrally resolved fluorescence data from intensity-based experiments. Another imaging method, fluorescence lifetime imaging microscopy (FLIM), is a widely used alternative to compute FRET efficiencies for each pixel in an image from the reduction of the fluorescence lifetime of the donors caused by FRET. In FLIM studies of oligomers with different proportions of donors and acceptors, the donor lifetimes may be obtained by fitting the temporally resolved fluorescence decay data with a predetermined number of exponential decay curves. However, this requires knowledge of the number and the relative arrangement of the fluorescent proteins in the sample, which is precisely the goal of FRET spectrometry, thus creating a conundrum that has prevented users of FLIM instruments from performing FRET spectrometry. Here, we describe an attempt to implement FRET spectrometry on temporally resolved fluorescence microscopes by using an integration-based method of computing the FRET efficiency from fluorescence decay curves. This method, which we dubbed time-integrated FRET (or tiFRET), was tested on oligomeric fluorescent protein constructs expressed in the cytoplasm of living cells. The present results show that tiFRET is a promising way of implementing FRET spectrometry and suggest potential instrument adjustments for increasing accuracy and resolution in this kind of study.
Full article
(This article belongs to the Section Molecular Biophysics)
►▼
Show Figures

Figure 1
Open AccessArticle
Antimicrobial Ionic Liquids: Ante-Mortem Mechanisms of Pathogenic EPEC and MRSA Examined by FTIR Spectroscopy
by
Patrick Mikuni-Mester, Christian Robben, Anna K. Witte, Kristina Linke, Monika Ehling-Schulz, Peter Rossmanith and Tom Grunert
Int. J. Mol. Sci. 2024, 25(9), 4705; https://doi.org/10.3390/ijms25094705 (registering DOI) - 26 Apr 2024
Abstract
Ionic liquids (ILs) have gained considerable attention due to their versatile and designable properties. ILs show great potential as antibacterial agents, but understanding the mechanism of attack on bacterial cells is essential to ensure the optimal design of IL-based biocides. The final aim
[...] Read more.
Ionic liquids (ILs) have gained considerable attention due to their versatile and designable properties. ILs show great potential as antibacterial agents, but understanding the mechanism of attack on bacterial cells is essential to ensure the optimal design of IL-based biocides. The final aim is to achieve maximum efficacy while minimising toxicity and preventing resistance development in target organisms. In this study, we examined a dose–response analysis of ILs’ antimicrobial activity against two pathogenic bacteria with different Gram types in terms of molecular responses on a cellular level using Fourier-transform infrared (FTIR) spectroscopy. In total, 18 ILs with different antimicrobial active motifs were evaluated on the Gram-negative enteropathogenic Escherichia coli (EPEC) and Gram-positive methicillin-resistant Staphylococcus aureus (MRSA). The results showed that most ILs impact bacterial proteins with increasing concentration but have a minimal effect on cellular membranes. Dose–response spectral analysis revealed a distinct ante-mortem response against certain ILs for MRSA but not for EPEC. We found that at sub-lethal concentrations, MRSA actively changed their membrane composition to counteract the damaging effect induced by the ILs. This suggests a new adaptive mechanism of Gram-positive bacteria against ILs and demonstrates the need for a better understanding before using such substances as novel antimicrobials.
Full article
(This article belongs to the Special Issue Molecular Mechanisms of Toxic and Activated Effects of Exogenous Compounds)
►▼
Show Figures

Figure 1
Open AccessArticle
Mixed-Lineage Leukaemia Gene Regulates Glucose-Sensitive Gene Expression and Insulin Secretion in Pancreatic Beta Cells
by
Satoshi Yoshino, Emi Ishida, Kazuhiko Horiguchi, Shunichi Matsumoto, Yasuyo Nakajima, Atsushi Ozawa, Masanobu Yamada and Eijiro Yamada
Int. J. Mol. Sci. 2024, 25(9), 4704; https://doi.org/10.3390/ijms25094704 (registering DOI) - 26 Apr 2024
Abstract
The escalating prevalence of diabetes mellitus underscores the need for a comprehensive understanding of pancreatic beta cell function. Interest in glucose effectiveness has prompted the exploration of novel regulatory factors. The myeloid/lymphoid or mixed-lineage leukaemia gene (MLL) is widely recognised for
[...] Read more.
The escalating prevalence of diabetes mellitus underscores the need for a comprehensive understanding of pancreatic beta cell function. Interest in glucose effectiveness has prompted the exploration of novel regulatory factors. The myeloid/lymphoid or mixed-lineage leukaemia gene (MLL) is widely recognised for its role in leukemogenesis and nuclear regulatory mechanisms through its histone methyltransferase activity in active chromatin. However, its function within pancreatic endocrine tissues remains elusive. Herein, we unveil a novel role of MLL in glucose metabolism and insulin secretion. MLL knockdown in βHC-9 pancreatic beta cells diminished insulin secretion in response to glucose loading, paralleled by the downregulation of the glucose-sensitive genes SLC2a1 and SLC2a2. Similar observations were made in MLL heterozygous knockout mice (MLL+/−), which exhibited impaired glucose tolerance and reduced insulin secretion without morphological anomalies in pancreatic endocrine cells. The reduction in insulin secretion was independent of changes in beta cell mass or insulin granule morphology, suggesting the regulatory role of MLL in glucose-sensitive gene expression. The current results suggest that MLL interacts with circadian-related complexes to modulate the expression of glucose transporter genes, thereby regulating glucose sensing and insulin secretion. Our findings shed light on insulin secretion control, providing potential avenues for therapeutics against diabetes.
Full article
(This article belongs to the Special Issue Steroids and Lipophilic Hormones, and Their Actions 3.0)
►▼
Show Figures

Figure 1
Open AccessReview
Autoimmunity, New Potential Biomarkers and the Thyroid Gland—The Perspective of Hashimoto’s Thyroiditis and Its Treatment
by
Ewa Tywanek, Agata Michalak, Joanna Świrska and Agnieszka Zwolak
Int. J. Mol. Sci. 2024, 25(9), 4703; https://doi.org/10.3390/ijms25094703 (registering DOI) - 26 Apr 2024
Abstract
Autoimmune thyroid disease (AITD) is the most common organic specific illness of the thyroid gland. It may manifest as the overproduction or the decline of thyroxine and triiodothyronine. Hyperthyroidism develops due to the overproduction of hormones as an answer to the presence of
[...] Read more.
Autoimmune thyroid disease (AITD) is the most common organic specific illness of the thyroid gland. It may manifest as the overproduction or the decline of thyroxine and triiodothyronine. Hyperthyroidism develops due to the overproduction of hormones as an answer to the presence of stimulatory antibodies against the TSH receptor. Hashimoto’s thyroiditis (HT) is generally characterized by the presence of thyroid peroxidase and thyroglobulin antibodies, with a concomitant infiltration of lymphocytes in the thyroid. Due to the progressive destruction of cells, AITD can lead to subclinical or overt hypothyroidism. Pathophysiology of AITD is extremely complicated and still not fully understood, with genetic, environmental and epigenetic factors involved in its development. Due to increasing incidence and social awareness of this pathology, there is an urgent need to expand the background concerning AITD. A growing body of evidence suggests possible ways of treatment apart from traditional approaches. Simultaneously, the role of potential new biomarkers in the diagnosis and monitoring of AITD has been highlighted recently, too. Therefore, we decided to review therapeutic trends in the course of AITD based on its pathophysiological mechanisms, mainly focusing on HT. Another aim was to summarize the state of knowledge regarding the role of new biomarkers in this condition.
Full article
(This article belongs to the Special Issue New Insights in Biomarkers of Autoimmune and Autoinflammatory Diseases)
►▼
Show Figures
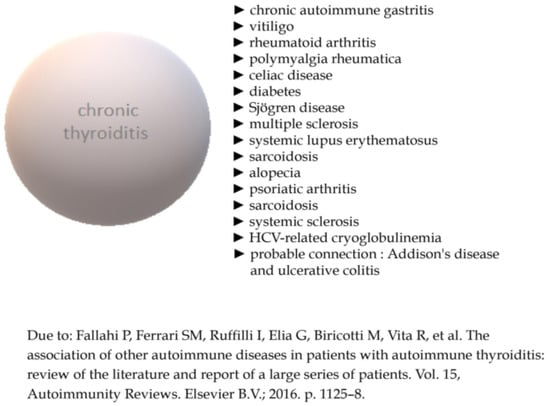

Figure 1
Open AccessArticle
Analysing the Cyanobacterial PipX Interaction Network Using NanoBiT Complementation in Synechococcus elongatus PCC7942
by
Carmen Jerez, Antonio Llop, Paloma Salinas, Sirine Bibak, Karl Forchhammer and Asunción Contreras
Int. J. Mol. Sci. 2024, 25(9), 4702; https://doi.org/10.3390/ijms25094702 (registering DOI) - 25 Apr 2024
Abstract
The conserved cyanobacterial protein PipX is part of a complex interaction network with regulators involved in essential processes that include metabolic homeostasis and ribosome assembly. Because PipX interactions depend on the relative levels of their different partners and of the effector molecules binding
[...] Read more.
The conserved cyanobacterial protein PipX is part of a complex interaction network with regulators involved in essential processes that include metabolic homeostasis and ribosome assembly. Because PipX interactions depend on the relative levels of their different partners and of the effector molecules binding to them, in vivo studies are required to understand the physiological significance and contribution of environmental factors to the regulation of PipX complexes. Here, we have used the NanoBiT complementation system to analyse the regulation of complex formation in Synechococcus elongatus PCC 7942 between PipX and each of its two best-characterized partners, PII and NtcA. Our results confirm previous in vitro analyses on the regulation of PipX-PII and PipX-NtcA complexes by 2-oxoglutarate and on the regulation of PipX-PII by the ATP/ADP ratio, showing the disruption of PipX-NtcA complexes due to increased levels of ADP-bound PII in Synechococcus elongatus. The demonstration of a positive role of PII on PipX-NtcA complexes during their initial response to nitrogen starvation or the impact of a PipX point mutation on the activity of PipX-PII and PipX-NtcA reporters are further indications of the sensitivity of the system. This study reveals additional regulatory complexities in the PipX interaction network, opening a path for future research on cyanobacteria.
Full article
(This article belongs to the Special Issue Advances in Protein-Protein Interactions 2.0)
Open AccessArticle
How to Personalize General Anesthesia—A Prospective Theoretical Approach to Conformational Changes of Halogenated Anesthetics in Fire Smoke Poisoning
by
Flavius Nicușor Truicu, Roni Octavian Damian, Mihai Alexandru Butoi, Vlad Ionuț Belghiru, Luciana Teodora Rotaru, Monica Puticiu and Renata Maria Văruț
Int. J. Mol. Sci. 2024, 25(9), 4701; https://doi.org/10.3390/ijms25094701 (registering DOI) - 25 Apr 2024
Abstract
Smoke intoxication is a central event in mass burn incidents, and toxic smoke acts at different levels of the body, blocking breathing and oxygenation. The majority of these patients require early induction of anesthesia to preserve vital functions. We studied the influence of
[...] Read more.
Smoke intoxication is a central event in mass burn incidents, and toxic smoke acts at different levels of the body, blocking breathing and oxygenation. The majority of these patients require early induction of anesthesia to preserve vital functions. We studied the influence of hemoglobin (HMG) and myoglobin (MGB) blockade by hydrochloric acid (HCl) in an interaction model with gaseous anesthetics using molecular docking techniques. In the next part of the study, molecular dynamics (MD) simulations were performed on the top-scoring ligand–receptor complexes to investigate the stability of the ligand–receptor complexes and the interactions between ligands and receptors in more detail. Through docking analysis, we observed that hemoglobin creates more stable complexes with anesthetic gases than myoglobin. Intoxication with gaseous hydrochloric acid produces conformational and binding energy changes of anesthetic gases to the substrate (both the pathway and the binding site), the most significant being recorded in the case of desflurane and sevoflurane, while for halothane and isoflurane, they remain unchanged. According to our theoretical model, the selection of anesthetic agents for patients affected by fire smoke containing hydrochloric acid is critical to ensure optimal anesthetic effects. In this regard, our model suggests that halothane and isoflurane are the most suitable choices for predicting the anesthetic effects in such patients when compared to sevoflurane and desflurane.
Full article
(This article belongs to the Special Issue Molecular Toxicology of New Drugs: New Insights)
Open AccessArticle
Genome-Wide Identification, Phylogenetic and Expression Analysis of Expansin Gene Family in Medicago sativa L
by
Yajing Li, Yangyang Zhang, Jing Cui, Xue Wang, Mingna Li, Lili Zhang and Junmei Kang
Int. J. Mol. Sci. 2024, 25(9), 4700; https://doi.org/10.3390/ijms25094700 (registering DOI) - 25 Apr 2024
Abstract
Expansins, a class of cell-wall-loosening proteins that regulate plant growth and stress resistance, have been studied in a variety of plant species. However, little is known about the Expansins present in alfalfa (Medicago sativa L.) due to the complexity of its tetraploidy.
[...] Read more.
Expansins, a class of cell-wall-loosening proteins that regulate plant growth and stress resistance, have been studied in a variety of plant species. However, little is known about the Expansins present in alfalfa (Medicago sativa L.) due to the complexity of its tetraploidy. Based on the alfalfa (cultivar “XinjiangDaye”) reference genome, we identified 168 Expansin members (MsEXPs). Phylogenetic analysis showed that MsEXPs consist of four subfamilies: MsEXPAs (123), MsEXPBs (25), MsEXLAs (2), and MsEXLBs (18). MsEXPAs, which account for 73.2% of MsEXPs, and are divided into twelve groups (EXPA-I–EXPA-XII). Of these, EXPA-XI members are specific to Medicago trunctula and alfalfa. Gene composition analysis revealed that the members of each individual subfamily shared a similar structure. Interestingly, about 56.3% of the cis-acting elements were predicted to be associated with abiotic stress, and the majority were MYB- and MYC-binding motifs, accounting for 33.9% and 36.0%, respectively. Our short-term treatment (≤24 h) with NaCl (200 mM) or PEG (polyethylene glycol, 15%) showed that the transcriptional levels of 12 MsEXPs in seedlings were significantly altered at the tested time point(s), indicating that MsEXPs are osmotic-responsive. These findings imply the potential functions of MsEXPs in alfalfa adaptation to high salinity and/or drought. Future studies on MsEXP expression profiles under long-term (>24 h) stress treatment would provide valuable information on their involvement in the response of alfalfa to abiotic stress.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
Open AccessReview
Anti-Obesity Therapeutic Targets Studied In Silico and In Vivo: A Systematic Review
by
Wendjilla F. de Medeiros, Ana Francisca T. Gomes, Ana Júlia F. C. Aguiar, Jaluza Luana C. de Queiroz, Ingrid Wilza L. Bezerra, Juliana Kelly da Silva-Maia, Grasiela Piuvezam and Ana Heloneida de A. Morais
Int. J. Mol. Sci. 2024, 25(9), 4699; https://doi.org/10.3390/ijms25094699 (registering DOI) - 25 Apr 2024
Abstract
In the age of information technology and the additional computational search tools and software available, this systematic review aimed to identify potential therapeutic targets for obesity, evaluated in silico and subsequently validated in vivo. The systematic review was initially guided by the research
[...] Read more.
In the age of information technology and the additional computational search tools and software available, this systematic review aimed to identify potential therapeutic targets for obesity, evaluated in silico and subsequently validated in vivo. The systematic review was initially guided by the research question “What therapeutic targets have been used in in silico analysis for the treatment of obesity?” and structured based on the acronym PECo (P, problem; E, exposure; Co, context). The systematic review protocol was formulated and registered in PROSPERO (CRD42022353808) in accordance with the Preferred Reporting Items Checklist for Systematic Review and Meta-Analysis Protocols (PRISMA-P), and the PRISMA was followed for the systematic review. The studies were selected according to the eligibility criteria, aligned with PECo, in the following databases: PubMed, ScienceDirect, Scopus, Web of Science, BVS, and EMBASE. The search strategy yielded 1142 articles, from which, based on the evaluation criteria, 12 were included in the systematic review. Only seven these articles allowed the identification of both in silico and in vivo reassessed therapeutic targets. Among these targets, five were exclusively experimental, one was exclusively theoretical, and one of the targets presented an experimental portion and a portion obtained by modeling. The predominant methodology used was molecular docking and the most studied target was Human Pancreatic Lipase (HPL) (n = 4). The lack of methodological details resulted in more than 50% of the papers being categorized with an “unclear risk of bias” across eight out of the eleven evaluated criteria. From the current systematic review, it seems evident that integrating in silico methodologies into studies of potential drug targets for the exploration of new therapeutic agents provides an important tool, given the ongoing challenges in controlling obesity.
Full article
(This article belongs to the Special Issue Anti-obesity Drug Discovery)
►▼
Show Figures
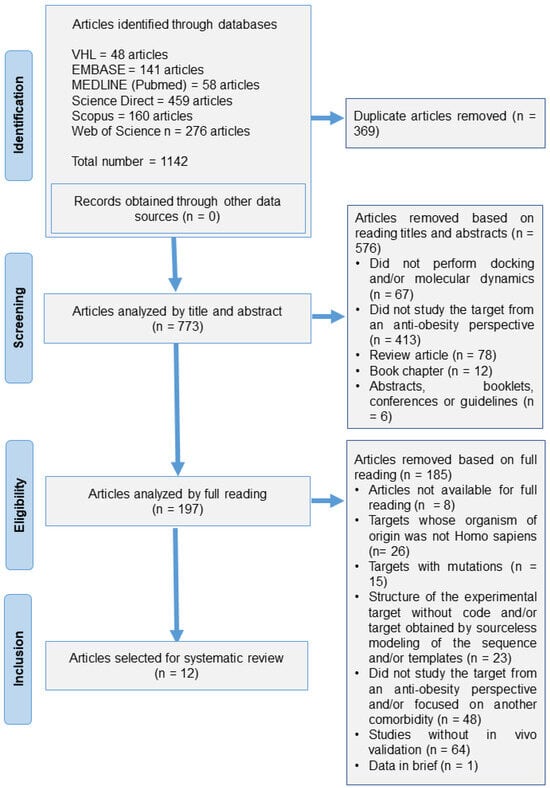
Figure 1
Open AccessArticle
Pure-Shift-Based Proton Magnetic Resonance Spectroscopy for High-Resolution Studies of Biological Samples
by
Haolin Zhan, Yulei Chen, Yinping Cui, Yunsong Zeng, Xiaozhen Feng, Chunhua Tan, Chengda Huang, Enping Lin, Yuqing Huang and Zhong Chen
Int. J. Mol. Sci. 2024, 25(9), 4698; https://doi.org/10.3390/ijms25094698 (registering DOI) - 25 Apr 2024
Abstract
Proton magnetic resonance spectroscopy (1H MRS) presents a powerful tool for revealing molecular-level metabolite information, complementary to the anatomical insight delivered by magnetic resonance imaging (MRI), thus playing a significant role in in vivo/in vitro biological studies. However, its further applications
[...] Read more.
Proton magnetic resonance spectroscopy (1H MRS) presents a powerful tool for revealing molecular-level metabolite information, complementary to the anatomical insight delivered by magnetic resonance imaging (MRI), thus playing a significant role in in vivo/in vitro biological studies. However, its further applications are generally confined by spectral congestion caused by numerous biological metabolites contained within the limited proton frequency range. Herein, we propose a pure-shift-based 1H localized MRS method as a proof of concept for high-resolution studies of biological samples. Benefitting from the spectral simplification from multiplets to singlet peaks, this method addresses the challenge of spectral congestion encountered in conventional MRS experiments and facilitates metabolite analysis from crowded NMR resonances. The performance of the proposed pure-shift 1H MRS method is demonstrated on different kinds of samples, including brain metabolite phantom and in vitro biological samples of intact pig brain tissue and grape tissue, using a 7.0 T animal MRI scanner. This proposed MRS method is readily implemented in common commercial NMR/MRI instruments because of its generally adopted pulse-sequence modules. Therefore, this study takes a meaningful step for MRS studies toward potential applications in metabolite analysis and disease diagnosis.
Full article
(This article belongs to the Special Issue Advances in Molecular and Cellular Imaging, Microscopy, and Biomedical Spectroscopy)
Open AccessReview
HIV-Associated Neurocognitive Disorder: A Look into Cellular and Molecular Pathology
by
Landon John-Patrick Thompson, Jessica Genovese, Zhenzi Hong, Meera Vir Singh and Vir Bahadur Singh
Int. J. Mol. Sci. 2024, 25(9), 4697; https://doi.org/10.3390/ijms25094697 (registering DOI) - 25 Apr 2024
Abstract
Despite combined antiretroviral therapy (cART) limiting HIV replication to undetectable levels in the blood, people living with HIV continue to experience HIV-associated neurocognitive disorder (HAND). HAND is associated with neurocognitive impairment, including motor impairment, and memory loss. HIV has been detected in the
[...] Read more.
Despite combined antiretroviral therapy (cART) limiting HIV replication to undetectable levels in the blood, people living with HIV continue to experience HIV-associated neurocognitive disorder (HAND). HAND is associated with neurocognitive impairment, including motor impairment, and memory loss. HIV has been detected in the brain within 8 days of estimated exposure and the mechanisms for this early entry are being actively studied. Once having entered into the central nervous system (CNS), HIV degrades the blood–brain barrier through the production of its gp120 and Tat proteins. These proteins are directly toxic to endothelial cells and neurons, and propagate inflammatory cytokines by the activation of immune cells and dysregulation of tight junction proteins. The BBB breakdown is associated with the progression of neurocognitive disease. One of the main hurdles for treatment for HAND is the latent pool of cells, which are insensitive to cART and prolong inflammation by harboring the provirus in long-lived cells that can reactivate, causing damage. Multiple strategies are being studied to combat the latent pool and HAND; however, clinically, these approaches have been insufficient and require further revisions. The goal of this paper is to aggregate the known mechanisms and challenges associated with HAND.
Full article
(This article belongs to the Special Issue Mechanisms of Neurotoxicity)

Journal Menu
► ▼ Journal Menu-
- IJMS Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal Browser-
arrow_forward_ios
Forthcoming issue
arrow_forward_ios Current issue - Vol. 25 (2024)
- Vol. 24 (2023)
- Vol. 23 (2022)
- Vol. 22 (2021)
- Vol. 21 (2020)
- Vol. 20 (2019)
- Vol. 19 (2018)
- Vol. 18 (2017)
- Vol. 17 (2016)
- Vol. 16 (2015)
- Vol. 15 (2014)
- Vol. 14 (2013)
- Vol. 13 (2012)
- Vol. 12 (2011)
- Vol. 11 (2010)
- Vol. 10 (2009)
- Vol. 9 (2008)
- Vol. 8 (2007)
- Vol. 7 (2006)
- Vol. 6 (2005)
- Vol. 5 (2004)
- Vol. 4 (2003)
- Vol. 3 (2002)
- Vol. 2 (2001)
- Vol. 1 (2000)
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomedicines, Cells, IJMS, Life, Oxygen
Oxidative Stress and Inflammation, 2nd Volume
Topic Editors: Mohamad Allaw, Ines Castangia, Maria Letizia Manca, Matteo Perra, Amparo NacherDeadline: 31 May 2024
Topic in
BioChem, Biomedicines, Biomolecules, IJMS, Metabolites, Molecules
Natural Products in Prevention and Therapy of Metabolic Syndrome
Topic Editors: Jianbo Wan, Ligen LinDeadline: 30 June 2024
Topic in
Cells, Diseases, Healthcare, IJMS, Vaccines
Inflammation: The Cause of all Diseases 2.0
Topic Editors: Vasso Apostolopoulos, Jack Feehan, Vivek P. ChavdaDeadline: 31 July 2024
Topic in
Biomedicines, CIMB, Endocrines, IJMS, JMP, Life, Reprod. Med.
Pathogenesis of Pregnancy-Related Complications 2.0
Topic Editors: Ilona Hromadnikova, Katerina KotlabovaDeadline: 31 August 2024

Conferences
Special Issues
Special Issue in
IJMS
Monoclonal Antibodies and Their Functional Fragments in Research, Diagnosis and Therapy 3.0
Guest Editors: Menotti Ruvo, Annamaria SandomenicoDeadline: 30 April 2024
Special Issue in
IJMS
Pharmacogenetics and Personalized Medicine 3.0
Guest Editors: José A. Riancho, Gloria RavegniniDeadline: 20 May 2024
Special Issue in
IJMS
Molecular Mechanisms of Angiogenesis and Cancer
Guest Editor: Vijay AvinDeadline: 30 May 2024
Topical Collections
Topical Collection in
IJMS
Feature Papers in Bioactives and Nutraceuticals
Collection Editor: Maurizio Battino
Topical Collection in
IJMS
State-of-the-Art Molecular Microbiology in Poland
Collection Editors: Alicja Wegrzyn, Satish Raina
Topical Collection in
IJMS
Computational, Structural and Spectroscopic Studies of Enzyme Mechanisms, Inhibition and Dynamics
Collection Editor: Christo Christov